

Protein and chitin preservation in polymeric sheets in Miocene Ecphora gardnerae shells from Maryland, USA
TIMOTHY P. CLELAND, G. ASHER NEWSOME, ERIN R. BIRDSALL, THOMAS LAM, JOHN R. NANCE, and ROBERT M. HAZEN
Cleland, T.P., Newsome, G.A., Birdsall, E.R., Lam, T., Nance, J.R., and Hazen, R.M. 2025. Protein and chitin preservation in polymeric sheets in Miocene Ecphora gardnerae shells from Maryland, USA. Acta Palaeontologica Polonica 70 (3): 507–516.
The shells of muricoid gastropod Ecphora gardnerae from the Miocene St. Marys Formation of Maryland, USA, have been shown to contain polymeric sheets within the calcareous matrix. These membranes were found to have amino acids and complex sugar compounds. To explore these biomolecular features further, four new E. gardnerae shells were collected and analyzed for proteins and sugar compounds. Using direct analysis in real time (DART) mass spectrometry, we were able to show that the membranes contain chitin. With mass spectrometry-based paleoproteomic techniques utilizing a hydroxylamine extraction or sample preparation by easy extraction and digestion (SPEED), we found preserved peptides from all four shells including peptides that are specific to Gastropoda or Mollusca. The membranes derive from the calcitic layer of the shell and are likely analogous to chitin membranes used by other extant mollusks to control nucleation, growth, and physical properties in the calcareous layers, but future research in gastropod biology will help to address the function of these membranes.
Key words: Gastropoda, Muricidae, Ecphora, paleoproteomics, DART-MS, chitin, Miocene, USA.
Timothy P. Cleland [clelandtp@si.edu; ORCID: https://orcid.org/0000-0001-9198-2828 ], G. Asher Newsome [newsomeg@si.edu; ORCID: https://orcid.org/0000-0003-1683-2197 ], and Thomas F. Lam [lamt@si.edu; ORCID: https://orcid.org/0000-0002-0908-0938 ], Museum Conservation Institute, Smithsonian Institution, Suitland, MD 20746 USA.
Erin R. Birdsall [birdsalle@si.edu; ORCID: https://orcid.org/0000-0002-1972-7475 ], Museum Conservation Institute, Smithsonian Institution, Suitland, MD 20746 USA. Smithsonian National Museum of the American Indian, Suitland, MD 20746, USA.
John R. Nance [John.Nance@calvertcountymd.gov; ORCID: https://orcid.org/0000-0003-1246-1237 ], Department of Paleontology, Calvert Marine Museum, PO Box 97, Solomons, MD, 20688, USA.
Robert M. Hazen [rhazen@carnegiescience.edu; ORCID: https://orcid.org/0000-0003-4163-8644 ], Earth and Planets Laboratory, Carnegie Science, Washington, DC 20015 USA.
Received 21 August 2024, accepted 23 July 2025, published online 10 September 2025.
Copyright © 2025 T.P. Cleland et al. This is an open-access article distributed under the terms of the Creative Commons Attribution License (for details please see http://creativecommons.org/licenses/by/4.0/), which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are credited.
Introduction
To date, paleoproteomics has focused primarily on vertebrate materials (e.g., Cleland et al. 2015, 2016; Presslee et al. 2018; Cappellini et al. 2019; Welker et al. 2019; Brown et al. 2021; Demarchi et al. 2022), particularly because vertebrates are better represented within protein databases than invertebrates. Despite the limited database availability, recent studies of invertebrate paleoproteins (Sakalauskaite et al. 2019, 2020) has yielded the ability to separate closely related mollusk species used in archaeological samples. Additionally, invertebrate shells have been understudied because they have a much lower organic content (1–5%) compared to vertebrate bone (Huang and Zhang 2022). Furthermore, invertebrate mineralization follows a different pathway than vertebrate mineralization and is predominantly extracellular controlling nucleation and formation of the CaCO3 (Huang and Zhang 2022).
Ecphora is an extinct genus of gastropods classified in Muricidae (Ward and Gilinsky 1988) with distinct red-orange coloration, prominent costae, and a thick external layer of calcite (Carter et al. 1994; Nance et al. 2015). Ecphora gardnerae is of special interest to the Calvert Cliffs area, Maryland, USA, because it retains its coloration while other mollusk shells in the same deposits are white chalky shells (Nance et al. 2015). In 2015, polymeric sheets derived from Ecphora gardnerae shells were described and were found to contain amino acids and evidence for complex sugar-amino acid reactions (Nance et al. 2015). Questions remained regarding what kind of proteins were contained within the polymeric sheets as well as the identity of the sugar compound within the sheets.
Since the original Ecphora membrane publication was published, development in the area of shell/invertebrate paleoproteomics/proteomics has greatly expanded. Protein methods to extract mineral associated and/or chitin associated proteins have been optimized (Tan et al. 2015; Sakalauskaite et al. 2020) to maximize the amount of protein information that can be used for species identification or biology. Additionally, application of paleoproteomics to fossils has resulted in the detection of biomineral proteins from Pleistocene coral (Drake et al. 2020). With the continued expansion of paleoproteomics from invertebrates, direct evaluation of biomineralization evolution or molecular phylogenetics of extinct species may be possible, similar to what has been done from vertebrate remains (e.g., Welker et al. 2015, 2019, 2020; Cappellini et al. 2019).
In this study, four Ecphora gardnerae shells were collected from the St. Marys Formation Miocene sediments in Maryland, USA, to biochemically characterize the membranes for preserved proteins and small molecule composition using proteomics and high-resolution mass spectrometry. We also discuss the preservation and taphonomic state of the shells.
Institutional abbreviations.—CMM, Calvert Marine Museum, Solomons, MD, USA.
Other abbreviations.—CDS, coding DNA sequence; Da, Dalton; G-PTM-D, Global-Post-translational Modification-Detection; i.d., internal diameter; H, histidine; L, liter; K, lysine; M, molar; μL, microliter; mL, milliliter; mW, milliwatt; mM millimolar; m/z, mass-to-charge ratio; nL, nanoliter; R, arginine; rcf, relative centrifugal force; W, tryptophan; Y, tyrosine.
Geological setting
The Miocene Calvert, Choptank, and St. Marys formations are well exposed throughout the Southern Maryland region of Calvert, Charles, and St. Mary’s Counties, Maryland, USA. The Calvert Cliffs represent the best outcrop of these formations; however, several smaller significant localities exist including Windmill Point, St. Mary’s County, Maryland, USA. The St. Marys Formation is the youngest of the three formations and is further divided into three members, Conoy Member, Little Cove Point Member, and Windmill Point Member (Ward and Andrews 2008). The Little Cove Point Member and the Windmill Point Member are both present at Windmill Point and samples were collected from each (Fig. 1). The Little Cove Point Member is a blue-gray very fine grained silty sand and the Windmill Point Member is an orange-brown fine grained sand (Ward and Andrews 2008).
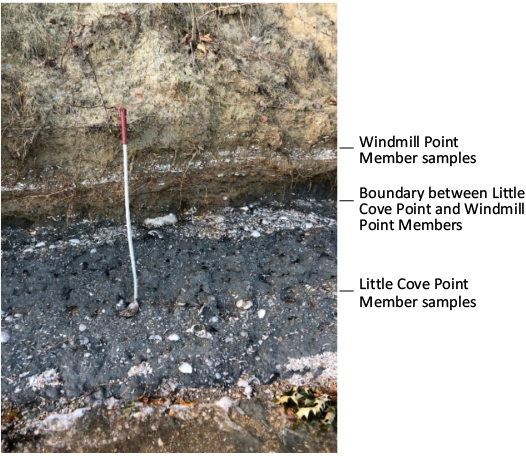

Fig. 1. Photograph of the collecting locality of the St. Marys Formation at Windmill Point, St. Mary’s County, Maryland, USA (GPS coordinates: 38°09’46.9”N 76°27’17.1”W). Collecting scoop is 107 cm for scale.
Material and methods
Samples.—Four shells of Ecphora gardnerae Wilson, 1987, were collected by JRN and RMH from the St. Marys Formation at Windmill Point, St. Mary’s County, Maryland, USA (GPS coordinates: 38°09’46.9”N 76°27’17.1”W) and are deposited in the Calvert Marine Museum (CMM) collection. The first two shells (CMM-I-6357, 6358) are ~10 Ma (Late Miocene) and were collected from Bed 23 of the Little Cove Point Member (Fig. 2). The second two shells (CMM-I-6359, 6360) are ~8 Ma (Late Miocene) and were collected from Bed 24, Windmill Point Member (Fig. 2). All shells were retained within their original matrix in aluminum foil and placed on indicator silica gel until arrival at the Museum Conservation Institute. TPC removed the sedimentary grains with aseptic techniques and photographed the shells. Additionally, two turritellid shells (one 8 Ma and one 10 Ma) found within the entombing sediments of the Ecphora were used for preservational comparison.
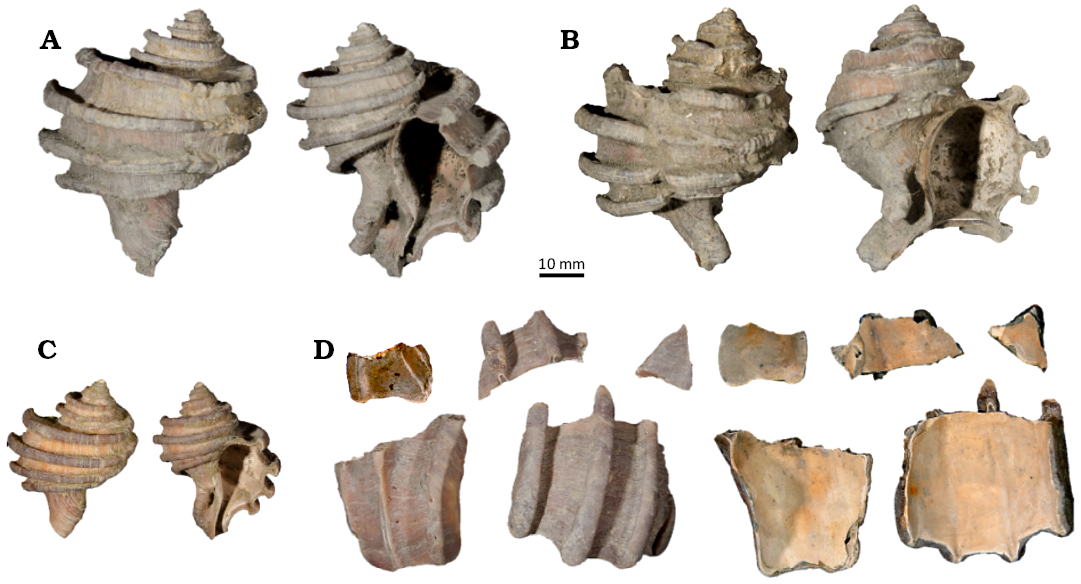

Fig. 2. Images of each shell of the muricoid gastropod Ecphora gardnerae Wilson, 1987 (St. Marys Formation, Miocene, Maryland, USA) used in this study. CMM-I-6357 (A) and CMM-I-6358 (B) are from Bed 23 (~10 Ma) of the Little Cove Point Member and CMM-I-6359 (C) and CMM-I-6360 (D) are from Bed 24 of the Windmill Point Member (~8 Ma).
Sample preparation for DART-MS.—Shell fragments from all four shells were taken with pliers from the body whorl adjacent to the outer lip of the aperture and demineralized in 1.2 M HCl until bubbling stopped and the polymeric sheet floated within the HCl column. Membranes were taken by pipette tip and transferred to new 1.5 mL tubes and washed with MilliQ water to neutralize the remaining HCl. After washing, membranes were centrifuged at 17 000 rcf for 2 min and the water was removed.
Direct Analysis in Real Time (DART) of molecules up to 600 Da was performed using a DART 100 probe with a simplified voltage and pressure (SVP) controller (IonSense). In the methodology, analytes are thermally desorbed off solid or semi-solid surfaces into the gas phase for ionization, chiefly via proton transfer from water clusters created as the post-plasma flow mixes with ambient air (Cody et al. 2005). The DART probe was positioned in front of an Orbitrap Elite mass spectrometer (Thermo Fisher Scientific) with a spacing of 7 mm between the ceramic cap of the DART and the transfer tube. Supplementary pumping (3.4 L/min) was provided by an IonSense Vapur flange fitted over the instrument inlet. Samples of polymeric sheet were held with methanol-cleaned fine tip tweezers in the gap between the DART ionization source and the transfer tube. The DART helium temperature was set to 350°C, and mass spectra were collected in positive ion mode and negative ion mode. Data were collected for short periods both before and after introducing the sample to the gas plume to collect background signal for comparison. Spectra were collected across a m/z 60–600 range at a resolving power of 120 000. Targeted fragmentation for chitin (C8H13NO5-H2O) was performed with collision induced dissociation (CID) at 30 collision energy (unitless).
Proteomic sample preparation.—Hydroxylamine extraction for chitin bound proteins (Tan et al. 2015): Shell fragments (19.4–28 mg) were taken from the body whorl adjacent to the outer lip of the aperture and demineralized with 1.2 M HCl until the polymeric sheets floated within the HCl in ThermoScientific Safelock microcentrifuge tubes. The HCl was removed carefully and then 2 M hydroxylamine, 2 M guanidine HCl , 0.2 M K2CO3, pH 9.0 (Tan et al. 2015) was added at a 10:1 ratio to the original mass of the shell or an empty tube (extraction blank) and incubated overnight shaking at 45°C. After incubation, remaining membrane was pelleted at 17 000 rcf for 2 min. 100 μL of protein extraction was digested using the single pot solid-phase sample preparation (SP3) technique (Moggridge et al. 2018; Cleland 2018). In short, proteins were bound to 10 μL of 1:1 hydrophobic:hydrophilic SeraMag beads (GE) for 10 minutes after the addition of 100% ethanol to a final concentration of 50%. The supernatant was removed after pulling the beads to a magnet then the beads were washed three times with 80% ethanol. After washing, the beads were resuspended in 100 μL of 50 mM ammonium bicarbonate and digested with 0.4 μg of Promega modified trypsin overnight at 37°C. Peptides were desalted and concentrated using Empore C18 stage tips (Rappsilber et al. 2007). Peptides were dried under vacuum at 45°C and then resuspended with 10 μL of 0.1% formic acid.
Sample preparation by easy extraction and digestion (SPEED) extraction (Doellinger et al. 2020): Shell fragments (17.7–42.8 mg) were demineralized in 1.2 M HCl until the sheets were free floating. After removal of the HCl, 100% trifluoroacetic acid (TFA) was added at a 4:1 ratio based on the original shell mass or an empty tube (extraction blank) and incubated for 10 min at room temperature. Subsequently, a 10-times volume of 2 M TrisBase was added to neutralize the TFA. Protein concentration was measured using turbidity at 360 nm against the Thermo Scientific Remel McFarland Equivalence Turbidity Standard #3. To 5 μg of each sample a 1:10 ratio (400 mM chloroacetamide/100 mM TCEP) was added and incubated for 5 min at 95°C. After reduction and alkylation, solution was diluted 5× with MilliQ water. 2 μg of Promega modified trypsin was added and incubated for 2 h at 37°C. Peptides were desalted and concentrated using Empore C18 stage tips (Rappsilber et al. 2007). Peptides were dried under vacuum at 45°C and then resuspended with 10 μL of 0.1% formic acid.
LC-MS/MS.—For the hydroxylamine samples, 7.5 μL of peptides or extraction blank were injected onto a ThermoScientific Acclaim PepMap 100 trap column (100 µm i.d. × 2 cm, 5 µm particle size) and separated on a ThermoScientific Acclaim PepMap RSLC analytical column (75 µm i.d. × 25 cm, 2 µm particle size) at 300 nL/min with the following gradient: 0–2 min 2% B, 2–68 min 2–31% B, 68–70 min 31–90% B, 70–73 min 90% B, 73–74 min 90–2% B, 74–90 min 2% B. For the SPEED samples, 1 μL of peptides or the buffer blank were injected onto a ThermoScientific Acclaim PepMap 100 trap columns (100 µm i.d. × 2 cm, 5 µm particle size) and separated on a ThermoScientific Acclaim PepMap RSLC analytical column (75 µm i.d. × 25 cm, 2 µm particle size) at 100 nL/min with the following gradient: 0–2 min 2% B, 2–68 min 2–31% B, 68–70 min 31–90% B, 70–73 min 90% B, 73–74 min 90–2% B, 74–120 min 2% B. Buffer A was 0.1% Optima formic acid (FisherScientific) in Optima water and Buffer B was 0.1% Optima Formic Acid in Optima acetonitrile. Data for both extractions were collected on a Thermo Scientific Orbitrap Elite as data dependent analysis (DDA) with the following parameters: MS1 was collected from 375–2000 m/z at 60 000 resolving power with 1E6 AGC and 100 ms max inject time. A top 10 method was used with the following parameters: 15 000 resolving power, 30% NCE HCD, default charge state 2, 0.1 ms activation time, 5 m/z isolation width, fixed first mass at 100 m/z, 5E5 AGC, and 250 ms max inject time. Between the SPEED samples, a 0.1% formic acid wash was included, but hydroxylamine samples were run sequentially.
Strombus and Conus genome conversion.—Genomic databases from Strombus pugilis (Fig. 3) (PRJNA655996, Sullivan et al. 2021; scaffold kindly provided by Alexis Sullivan), Conus ventricosus (ASM1839881v2; downloaded from the National Center for Biotechnology Information March 31, 2025), Conus betulinus (ASM1680195v1; downloaded from the National Center for Biotechnology Information March 31, 2025), Conus consors (ASM419361v1; downloaded from the National Center for Biotechnology Information March 31, 2025), and Conus tribblei (ASM126257v1; downloaded from the National Center for Biotechnology Information March 31, 2025) were annotated using GeMoMa v. 1.9 (Keilwagen et al. 2016, 2018) comparing against the following proteomes: Lottia gigantea, Pomacea canaliculata, Elysia chlorotica, Candidula unifasciata. GeMoMa is a homology-based gene prediction software that uses reference genomes to infer the annotations in a target genome. Following the annotation pipeline, annotated proteins and CDS sequences were extracted from the genomic FASTA file using the GeMoMa Extractor tool.

Fig. 3. General phylogenetic tree of Caenogastropoda (adapted from Ponder et al. 2008). Ecphora falls within Muricidae (magenta), Conus falls within Conidae (yellow), and Strombus falls within Strombidae (blue).
Mass Spectrometry Data Analysis.—Manual analysis of the most abundant ions observed with DART-MS for both the precursor and CID fragment masses were performed for both positive and negative mode ions. N-acetylglucosamine (GlcNAc) oxonium ions were annotated following the masses in Halim et al. (2014).
For the proteomic analyses, a Uniprot Gastropoda database (downloaded November 11, 2022) or the above Strombus pugilis (created April 26, 2023) or Conus species (created April 7, 2025) databases and the default MetaMorpheus contaminants database were used. All proteomic data were searched with MetaMorpheus 1.0.2 or 1.0.5. For the calibration task, default settings were used. For the G-PTM-D task, default settings were used with all modifications from Common Biological, Less Common, and Common Artifact. A custom modification list of diagenetic modifications was also used (imidazolone on R, methyl-imidazolone on R, aspartic acid from H, dioxidation of W, dioxidation of Y, dioxidation on H, hydroxyglutamic acid from H, aminoadipic acid on K, carbamyl on K, carboxyethylamine on K, carboxymethyl on K, ornithine on R, oxidation on W, oxidation on Y, oxidation on H, tryptophan to kynurenine, tryptophan to oxolactone). Either trypsin or a hydroxylamine-trypsin protease was used for the search task with default settings and generate complementary ions on. All peptide spectral matches (PSMs), peptides, and proteins were filtered at 1% false discovery rate and a Score ≥ 9. Peptide sequences were finally filtered based on the extraction blank files (for the list of peptides see SOM, Supplementary Online Material available at http://app.pan.pl/SOM/app70-Cleland_etal_SOM.pdf).
Peptide analysis.—PSM files were combined with RStudio v. RStudio 2023.03.0+386 and a unique list of peptides was output for Basic Local Alignment Search Tool (BLASTp) searching on NCBI. Peptides were evaluated for if they matched Gastropoda, other Mollusca, or other taxa (e.g., bacteria, mammals, fungi). Protein classification was performed with Panther 19.0. The Panther Classification System organizes proteins by their function, biological location, and biological pathway (Thomas et al. 2022).
Raman microscopy.—A Thermo Scientific DXR3 microscope was used to evaluate the mineral composition on the inner and outer surfaces of each sample with 633 nm laser excitation with 5 mW laser power and a spatial resolution of about 1 µm, focused through either a 50× (CMM-I-6357 inner, CMM-I-6358 inner) or 100× (CMM-I-6357 outer, CMM-I-6358 outer, CMM-I-6359 inner, CMM-I-6359 outer, CMM-I-6360 inner, CMM-I-6360 outer) objective. Objective selection was based on sample constraints. Spectra were collected for 7 seconds over 25 accumulations after 40 background accumulations. Raman spectra were collected, examined, and compared to literature results (Taylor et al. 2008) for identification. Peaks were identified using the Omnic Software (Thermo Fisher) “Find Peaks” function over the 100 cm-1 to 1100 cm-1 region. Fluorescent background was corrected using asymmetrically reweighted penalized least-squares smoothing (Baek et al. 2015). Photographs of samples were taken with Omnic Atlµs software (Thermo Scientific) before analysis.
In addition to the Ecphora shells, two turritellid shells (one 8 Ma, Late Miocene and one 10 Ma, Late Miocene) that were found within the entombing sediment were also analyzed using the same parameters at 50× magnification.
Scanning Electron Microscopy.—HCl demineralized, HCl etched (i.e., half of the shell fragments were incubated in 1.2 M HCl until membranes became visible on the surface then the samples were washed 3 times with MilliQ water), SPEED extracted, and hydroxylamine extracted membranes were washed three times with 1 mL of MilliQ water to neutralize and dilute the salts. Samples were adhered onto an aluminum stub and scanning electron microscopy (SEM) in the modes of secondary electron mode and backscattered electron (BSE) mode was performed at 15kV using a JEOL JSM-IT710HR in variable pressure. Images were measured with ImageJ 1.53t.
Minimization of contamination.—All shells studied here were collected and retained within their original sediments while stored within aluminum foil and indicator silica gel. They were prepared by TPC and photographed. All prepared shells were stored within the original aluminum foil until sampling with methanol cleaned pliers. Because of our focus on only the mineral contained chitin-protein membranes, we did not apply additional washing to the external surfaces of the shell (e.g., with bleach). The membranes exposed during the demineralization process were only handled by pipette tip. Additionally, no modern gastropod shells have ever been studied in our spaces, so contamination with modern gastropod proteins is very unlikely.
Results
Reflecting the original study (Nance et al. 2015), we were able to find polymeric sheets within the shells of newly collected E. gardnerae shells. Below, we discuss the shell preservation state and characterization of the membranes external to the mineralized shell.
Evaluation of inorganic preservation.—To support that the polymeric membranes derive from the E. gardnerae shells, we evaluated the preservation of the mineral using Raman spectroscopy and SEM. We evaluated the preservation of the mineral phase of the shells with Raman spectroscopy. Calcite and aragonite are able to be distinguished by peaks between 100–1100 cm-1 (calcite: 155 cm-1, 282 cm-1, 711 cm-1, 1085 cm-1; aragonite: 152 cm-1, 206 cm-1, 705 cm-1, 1085 cm-1) (Taylor et al. 2008, DeCarlo 2018). Across the samples, four peaks were observed in spectra for the external surface of the shell at 155 cm-1, 281 cm-1, 714 cm-1, and 1086 cm-1 consistent with calcite (Fig. 4, SOM: table S5, figs. S1–S3). Four major peaks are similarly observed in spectra of the inner surface, though at slightly different frequencies: 152 cm-1, 206 cm-1, 703 cm-1, and 1085 cm-1. These peak centers are consistent with aragonite.
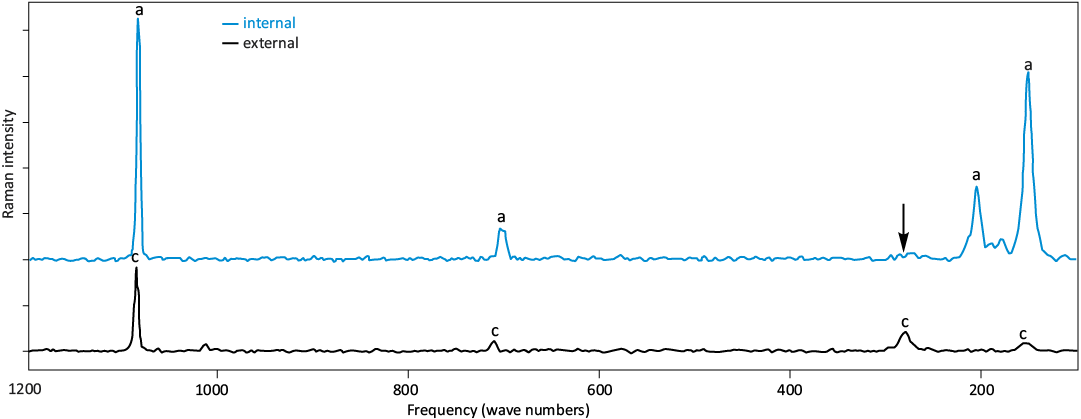
Fig. 4. Muricoid gastropod Ecphora gardnerae (CMM-I-6358; Bed 23 Little Cove Point Member of the St. Marys Formation, Miocene, Maryland, USA) Raman spectrum of internal (blue) and external (black) surfaces. a is peaks for aragonite, c is calcite. Black arrow indicates possible calcite peak on the internal surface.
Under light microscopy within the Raman microscope, the aragonite surface (SOM: fig. S4A) looks different from the calcite surface (SOM: fig. S4B). Using SEM, we were able to visualize the internal and external surfaces of each shell. We can clearly see calcite rhomboids (SOM: fig. S8) and needles of aragonite on the internal surfaces (SOM: fig. S8). Additionally, the crystal size reflects the relative maturity of the shells with smaller crystals (SOM: fig. S8) present in the smallest shell (CMM-I-6359) compared to larger crystals present in larger shells (SOM: fig. S8)
Characterization of the polymeric membranes.—The polymeric sheets derived from the new E. gardnerae shells examined in this study were found to be composed of chitin and proteins with additional biomolecule and diagenetic products present. To fully characterize them, we localized them to the external calcitic layer of the shells using SEM (SOM: fig. S8). After etching the shell, the membranes become visible in the orange-colored part of the shell and are not derived from the crossed-lamellar layer (SOM: fig. S9). Prior to the partial demineralization, membranes are not visible from the external surface (SOM: fig. S8) suggesting that they are fully contained within the calcite layers, but based on sheet structure likely are intercrystalline. Reflecting this location, these intact membranes are released only during the demineralization process (SOM: fig. S10), reducing access to external diagenetic processes (e.g., microbial interaction). Using DART-MS/MS, we were able to find chitin (as C8H13NO5-H2O; [M+H]: m/z 204.0860; [M-H]: m/z 202.0720) and its fragmentation products (Fig. 5) from all four shells. Urea was detected in CMM-I-6359 and CMM-I-6358, which is likely a diagenetic product of protein breakdown (Mackie et al. 2018).
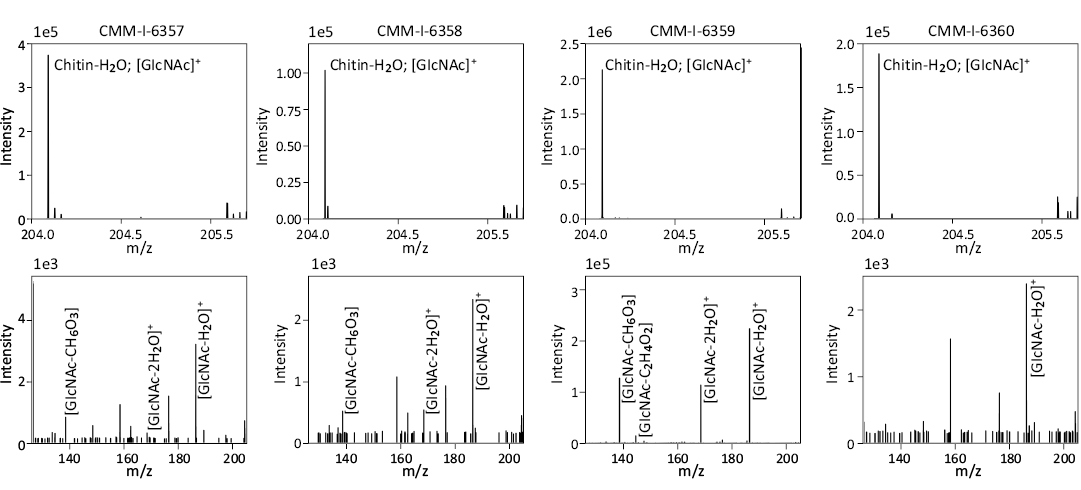
Fig. 5. Muricoid gastropod Ecphora gardnerae (St. Marys Formation, Miocene, Maryland, USA) chitin spectra from each membrane. Top row: Precursor masses for chitin minus water. Bottom row: Collision induced dissociation spectra of chitin mass-to-charge ratio (m/z) 204.0860. GlcNAc = N-acetylglucosamine. CMM-I-6357 and CMM-I-6358 are from Bed 23 (~10 Ma) of the Little Cove Point Member and CMM-I-6359 and CMM-I-6360 are from Bed 24 of the Windmill Point Member (~8 Ma).
Because the sheets were composed of chitin, we applied an extraction method designed for extracting proteins from chitin (i.e., the hydroxylamine method). With this method, we found a total of 222 unique peptides (468 PSMs). Specifically, we found 22 unique peptides (48 PSMs; SOM: table S1) from the Strombus database, 35 unique peptides (67 PSMs; SOM: table S1) from the Gastropoda database, 52 unique peptides (123 PSMs; SOM: table S1) from the Conus betulinus database, 36 unique peptides (80 PSMs; SOM: table S1) from the Conus consors database, 50 unique peptides (99 PSMs; SOM: table S1) from the Conus tribblei database, and 27 unique peptides (51 PSMs; SOM: table S1) from the C. betulinus database. To complement the hydroxylamine method, the SPEED approach was used because it has been shown to have high protein yields in a short period of time (Doellinger et al. 2020). We combined the high extraction yield with a reduced reduced flow, nanoLC flow rate to increase sensitivity (Brunner et al. 2022) to maximize the number of identifications. With this method, we found a total of 81 unique peptides (123 PSMs). Specifically, we found 6 unique peptides (10 PSMs; SOM: table S1) from the Strombus database, 19 unique peptides (32 PSMs; SOM: table S1) from the Gastropoda database, 16 unique peptides (24 PSMs; SOM: table S1) from the C. betulinus database, 8 unique peptides (13 PSMs; SOM: table S1) from the C. consors database, 11 unique peptides (18 PSMs; SOM: table S1) from the C. tribblei database, and 21 unique peptides (26 PSMs; SOM: table S1) from the C. betulinus database. For both types, 201 (413 PSMs; SOM: table S1) correspond to unique Gastropoda peptides, 5 (9 PSMs; Table S1) to unique Mollusca peptides, and 65 (169 PSMs; SOM: table S1) overlap with other species (e.g., bacteria, mammals). These overlapping sequences could derive from E. gardnerae because these membranes are trapped within the calcite matrix and not exposed until the demineralization process, but it is impossible to confirm because of the possibilities of normal laboratory contamination. In contrast to the overlapping sequences, the gastropod specific peptides more confidently identify that these peptides as deriving from E. gardnerae. Sample CMM-I-6359 from Bed 24 (~8 Ma) showed the best preservation for both extractions with 39 unique peptides (66 PSMs; Table S1). Sample CMM-I-6360 from Bed 24 had 23 unique peptides (35 PSMs; SOM: table S1), sample CMM-I-6357 had 24 unique peptides (40 PSMs; SOM: table S1), and CMM-I-6358 had 8 unique peptides (13 PSMs; SOM: table S1). Despite CMM-I-6359 being the smallest shell of the set, qualitatively it is the best preserved (i.e., the orange color is most distinct, especially compared to the bed 23 shells; Fig. 2) and that level of preservation is reflected in the protein and chitin preservation. The two extraction blanks were used to filter sequences that are the result of laboratory contamination. After using the FDR, score, and sequence filtering, no peptides remained in the extraction blank datasets.
Because no genome is available for E. gardnerae, we compared the peptide length to the entire identified protein to generate a sequence coverage value. Sequence coverage of the identified proteins (SOM: table S2) ranged from as little as 0.1% for hydroxylamine extractions from CMM-I-6359 and CMM-I-6360 to 68.9% for the SPEED extraction from CMM-I-6358. For all of the shells and extractions, the average sequence coverage was 4.5%.
We evaluated the origin of the various proteins (SOM: table S4) based on if a gene code could be determined for each one using the Panther database. We were able to evaluate 91 of the total 220 Gastropoda-only protein groups and found that 47 were cellular anatomical entities. Examining the membranes from CMM-I-6360 (SOM: fig. S5) after HCl demineralization, SPEED extraction, or hydroxylamine extraction shows several areas that are ~10–12 μm in diameter (SOM: fig. S6 yellow arrows) compared to the ~1 μm thick membrane itself. These larger areas may be the cellular components that we are extracting because they are not evident in the hydroxylamine extracted membranes. The one caveat being that despite dilution and washing, the hydroxylamine still precipitated crystals on the surface of the membranes (spherical crystals in SOM: fig. S5). A similar ~12 μm area is visible on membranes from CMM-I-6357 (SOM: fig. S7), but the membranes are rolled up too much to clearly differentiate exactly where these possible cellular areas may be. Membranes for the other 2 shells were also rolled up similarly to CMM-I-6357 (data not shown).
Discussion
Mineral preservation.—The combination of Raman signal and morphological patterning suggests that the mineral and shell preservation is good and reflects the original shell composition without impact from exogenous microbial organisms. Despite the overall good preservation, we observe that in samples CMM-I-6358 and CMM-I-6359, there may be some of the calcite 281 cm-1 peak present, though at low amounts (Fig. 4) suggesting some aragonite may have transformed into calcite. Consistent with the shell color (Fig. 2), CMM-I-6359 (SOM: fig. S2) shows some of the best preservation.
Membrane preservation.—The abundance of cellular components with other uncharacterized proteins (SOM: table S1) suggests that the membranes are cellular in origin with the cells on or within a chitin (Fig. 5) network. This refines the original description of the membranes as being a saccharide and protein, based on nuclear magnetic resonance and amino acid analysis (Nance et al. 2015). We were able to directly measure and fragment the intact chitin finding the diagnostic peaks representative of acetylglucosamine.
For the protein component, we found a variety of cellular and uncharacterized proteins. Unfortunately, gastropods have limited protein annotation, so many of our detected peptides remain from unknown proteins. With additional annotations of modern gastropod proteins, the annotations of these E. gardnerae peptides will be refined. We found relatively low levels of sequence coverage reflecting the age of the samples and the limited search space/lack of specific database for this species. Similar database issues/limitations were found for archaeological buttons (Sakalauskaite et al. 2019), where identification of species was limited by a very small number of molluscan representatives.
We detected a total of 22 gastropod-only peptides from shells from both the Little Cove Point Member (10 Ma) and Windmill Point Member (8 Ma) of the St. Marys Formation. These peptides derived from a variety of proteins including whirlin, cartilage matrix protein-like, nucleolin-like, and others (SOM: table S6). The overlapping sequences support the endogeneity of these peptide sequences because they are derived from multiple fossils from two localities. Additionally, the overlapping sequences reflect the protein sequence retention in E. gardnerae across 2 Ma of evolutionary time.
Relatively low levels of deamidation (SOM: table S3) and oxidation (Fig. 6) were found for the detected peptides showing that they are well preserved (Schroeter and Cleland 2016). Additionally, the low levels of deamidation may be the result of N-terminal asparagines and glutamines or post-translationally modified residues impeding the deamidation process. Asparagine is the N-terminal amino acid on 36 peptides and the second on 40 (combined 14.2% of the total N residues detected). Glutamine is N-terminal on 60 and the second on 23 as well (combined 15.3% of the total Q residues detected). Thirty glutamine residues were methylated (5.5% of the total Q residues detected). The location and post-translational modifications can impact the overall deamidation signal present and needs to be considered in all paleoproteomic studies. The limited damage may further be the result of encapsulation of the protein within the calcite mineral limiting access of both water and microbes to the organic molecules within. The association between the protein and chitin may also impact the preservation by stabilizing the local microenvironment in which the protein-chitin exist (Stankiewicz et al. 1997; Flannery et al. 2001; Montroni et al. 2021).
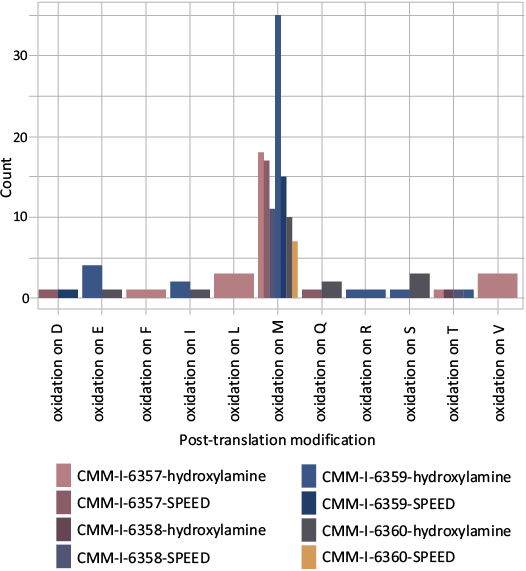
Fig. 6. Amino acid oxidation counts by sample CMM-I-6357 and CMM-I-6358 are from Bed 23 (~10 Ma) of the Little Cove Point Member and CMM-I-6359 and CMM-I-6360 are from Bed 24 of the Windmill Point Member (~8 Ma) of the St. Marys Formation, Miocene, Maryland, USA.
Impacts of membrane preservation on Ecphora gardnerae shells.—We hypothesize that the chitin-protein membrane preservation resulted in the excellent mineral preservation of all four E. gardnerae shells analyzed here. Unlike other gastropod shells from the same deposits that are composed exclusively of easily broken white aragonite (SOM: fig. S5), the relatively hard E. gardnerae shells possess the aragonitic crossed-lamellar layer retaining the original mineral size and orientation (SOM: fig. S8) and the distinct orange-red colored calcite layer where the membranes we characterized derive from. Based on the location within the mineralized shell itself, we suggest that these membranes within the calcitic layer are likely analogous to chitin membranes used by other extant mollusks to control nucleation, growth, and physical properties in the calcitic layers (De Paula and Silveira 2009; Checa et al. 2016). Exploration of extant gastropod shells for similar membranes will be necessary to fully understand their function. Because other extant mollusk possess chitin membranes within the aragonitic layer, we speculate that there may be filamentous (De Paula and Silveira 2009) or other interlamellar membranes that may not be darkly colored like the ones we observe here leading to the preservation of both shell layers. The inclusion of these darkly colored membranes within the calcitic layer of the shell, possible mineral controlling properties, and presence of the relatively insoluble chitin increased their chances of preservation in the E. gardnerae shells from both the Little Cove Point Member and Windmill Point Member of the St. Marys Formation.
Conclusions
We found that these E. gardnerae shells are well preserved based on their mineral content and morphological appearance supporting that these polymeric membranes derive from them allowing us to confidently show peptide and chitin preservation as old as 10 Ma. The cellular-chitin membranes within the calcitic layer are likely analogous to chitin membranes used by other extant mollusks to control nucleation, growth, and physical properties in the calcitic layers (De Paula and Silveira 2009; Checa et al. 2016), but future research in gastropod biology will help to address the function of these membranes.
Authors’ contributions
TC, conceptualization, formal analysis, methodology, writing: original draft, review, and editing; AN, formal analysis, methodology, writing: review and editing; EB, formal analysis, methodology, writing: review and editing; TL, formal analysis, methodology, writing: review and editing; JN, conceptualization, writing: original draft, review, and editing, collecting the specimens; RH, writing: review and editing, collecting the specimens.
Data availability
All RAW proteomics data, DART-MS data, and Meta Morpheus searches are available on Smithsonian Figshare at 10.25573/data.26513353, 10.25573/data.28924010, or 10.25573/ data.26513290
Acknowledgements
We thank Robert G. Jenkins (University of Kanazawa, Japan) and two anonymous referees for their thoughtful reviews. Funding for this project was provided by the Smithsonian Museum Conservation Institute Federal and Trust Funds to TPC, GAN, ERB, and TFL. Additional funding was provided in part by the citizens of Calvert County Maryland, the County Board of Calvert County Commissioners, and the Clarissa and Lincoln Dryden Endowment for paleontology at the Calvert Marine Museum. ERB was additionally funded by the Collections Care Initiative of the Smithsonian National Collections Program.
Editor: Andrzej Kaim
References
Baek, S.J., Park, A., Ahn, Y.J., and Choo, J. 2015. Baseline correction using asymmetrically reweighted penalized least squares smoothing. Analyst 140: 250–257. Crossref
Brown, S., Kozlikin, M., Shunkov, M., Derevianko, A., Higham, T., Douka, K., and Richter, K.K. 2021. Examining collagen preservation through glutamine deamidation at Denisova Cave. Journal of Archaeological Science 133: 105454. Crossref
Brunner, A.D., Thielert, M., Vasilopoulou, C., Ammar, C., Coscia, F., Mund, A., Hoerning, O.B., Bache, N., Apalategui, A., Lubeck, M., Richter, S., Fischer, D.S., Raether, O., Park, M.A., Meier, F., Theis, F.J., and Mann, M. 2022. Ultra-high sensitivity mass spectrometry quantifies single-cell proteome changes upon perturbation. Molecular Systems Biology 18 (3): e10798. Crossref
Cappellini, E., Welker, F., Pandolfi, L., Ramos-Madrigal, J., Samodova, D., Ruther, P.L., Fotakis, A.K., Lyon, D., Moreno-Mayar, J.V., Bukhsianidze, M., Jersie-Christensen, R.R., Mackie, M., Ginolhac, A., Ferring, R., Tappen, M., Palkopoulou, E., Dickinson, M.R., Stafford, T.W., Chan, Y.L., Gotherstrom, A., Nathan, S.K.S.S., Heintzman, P.D., Kapp, J.D., Kirillova, I., Moodley, Y., Agusti, J., Kahlke, R.D., Kiladze, G., Martinez-Navarro, B., Liu, S.L., Velasco, M.S., Sinding, M.H.S., Kelstrup, C.D., Allentoft, M.E., Orlando, L., Penkman, K., Shapiro, B., Rook, L., Dalen, L., Gilbert, M.T.P., Olsen, J.V., Lordkipanidze, D., and Willerslev, E. 2019. Early Pleistocene enamel proteome from Dmanisi resolves Stephanorhinus phylogeny. Nature 574: 103–107. Crossref
Carter, J.G., Rossbach, T.J., Robertson, K.J., and Ward, L.W. 1994. Morphological and microstructural evidence for the origin and early evolution of Ecphora (Mollusca: Gastropoda). Journal of Paleontology 68: 905–907. Crossref
Checa, A.G., Macias-Sanchez, E., Harper, E.M., and Cartwright, J.H. 2016. Organic membranes determine the pattern of the columnar prismatic layer of mollusc shells. Proceedings of the Royal Society B: Biological Sciences 283: 20160032. Crossref
Cleland, T.P. 2018. Human bone paleoproteomics utilizing the Single-Pot, Solid-Phase-Enhanced Sample Preparation method to maximize detected proteins and reduce humics. Journal of Proteome Research 17: 3976–3983. Crossref
Cleland, T.P., Schroeter, E.R., and Schweitzer, M.H. 2015. Biologically and diagenetically derived peptide modifications in moa collagens. Proceedings of the Royal Society B: Biological Sciences 282 (1808): 20150015. Crossref
Cleland, T.P., Schroeter, E.R., Feranec, R.S., and Vashishth, D. 2016. Peptide sequences from the first Castoroides ohioensis skull and the utility of old museum collections for palaeoproteomics. Proceedings of the Royal Society of London B: Biological Sciences 283: 20160593. Crossref
Cody, R.B., Laramée, J.A., and Durst, H.D. 2005. Versatile new ion source for the analysis of materials in open air under ambient conditions. Analytical Chemistry 77: 2297–2302. Crossref
De Paula, S.M. and Silveira, M. 2009. Studies on molluscan shells: contributions from microscopic and analytical methods. Micron 40: 669–90. Crossref
DeCarlo, T.M. 2018. Characterizing coral skeleton mineralogy with Raman spectroscopy. Nature Communications 9: 5325. Crossref
Demarchi, B., Mackie, M., Li, Z.H., Deng, T., Collins, M.J., and Clarke, J. 2022. Survival of mineral-bound peptides into the Miocene. Elife 11: e82849. Crossref
Doellinger, J., Schneider, A., Hoeller, M., and Lasch, P. 2020. Sample Preparation by Easy Extraction and Digestion (SPEED)—a universal, rapid, and detergent-free protocol for proteomics based on acid extraction. Molecular and Cellular Proteomics 19: 209–222. Crossref
Drake, J.L., Whitelegge, J.P., and Jacobs, D.K. 2020. First sequencing of ancient coral skeletal proteins. Scientific Reports 10: 19407. Crossref
Flannery, M.B., Stott, A.W., Briggs, D.E.G., and Evershed, R.P. 2001. Chitin in the fossil record: identification and quantification of D-glucosamine. Organic Geochemistry 32: 745–754. Crossref
Halim, A., Westerlind, U., Pett, C., Schorlemer, M., Rüetschi, U., Brinkmalm, G., Sihlbom, C., Lengqvist, J., Larson, G., and Nilsson, J. 2014. Assignment of saccharide identities through analysis of oxonium ion fragmentation profiles in LC-MS/MS of glycopeptides. Journal of Proteome Research 13: 6024–6032. Crossref
Huang, J.L. and Zhang, R.Q. 2022. The mineralization of molluscan shells: some unsolved problems and special considerations. Frontiers in Marine Science 9: 874534. Crossref
Keilwagen, J., Hartung, F., Paulini, M., Twardziok, S.O., and Grau, J. 2018. Combining RNA-seq data and homology-based gene prediction for plants, animals and fungi. BMC Bioinformatics 19: 189. Crossref
Keilwagen, J., Wenk, M., Erickson, J.L., Schattat, M.H., Grau, J., and Hartung, F. 2016. Using intron position conservation for homology-based gene prediction. Nucleic Acids Research 44: e89. Crossref
Mackie, M., Rüther, P., Samodova, D., Di Gianvincenzo, F., Granzotto, C., Lyon, D., Peggie, D.A., Howard, H., Harrison, L., Jensen, L.J., Olsen, J.V., and Cappellini, E. 2018. Palaeoproteomic Profiling of Conservation Layers on a 14th Century Italian Wall Painting. Angewandte Chemie International Edition 57: 7369–7374. Crossref
Moggridge, S., Sorensen, P.H., Morin, G.B., and Hughes, C.S. 2018. Extending the compatibility of the SP3 paramagnetic bead processing approach for proteomics. Journal of Proteome Research 17: 1730–1740. Crossref
Montroni, D., Sparla, F., Fermani, S., and Falini, G. 2021. Influence of proteins on mechanical properties of a natural chitin-protein composite. Acta Biomaterialia 120: 81–90. Crossref
Nance, J.R., Armstrong, J.T., Cody, G.D., Fogel, M.L., and Hazen, R.M. 2015. Preserved macroscopic polymeric sheets of shell-binding protein in the Middle Miocene (8 to 18 Ma) gastropod Ecphora. Geochemical Perspectives Letters 1: 1–9. Crossref
Ponder, W.F., Colgan, D.J., Healy, J., Nützel, A., Simone, L.R., and Strong, E.E. 2008. Caenogastropod phylogeny. In: W.F. Ponder and D.L. Lindberg (eds.), Molluscan Phylogeny, 331–383. U. California Press, Berkeley. Crossref
Presslee, S., Wilson, J., Woolley, J., Best, J., Russell, D., Radini, A., Fischer, R., Kessler, B., Boano, R., Collins, M., and Demarchi, B. 2018. The identification of archaeological eggshell using peptide markers. STAR: Science & Technology of Archaeological Research 3 (1): 89–99. Crossref
Rappsilber, J., Mann, M., and Ishihama, Y. 2007. Protocol for micro-purification, enrichment, pre-fractionation and storage of peptides for proteomics using StageTips. Nature Protocols 2: 1896–1906. Crossref
Sakalauskaite, J., Andersen, S.H., Biagi, P., Borrello, M.A., Cocquerez, T., Colonese, A.C., Dal Bello, F., Girod, A., Heumuller, M., Koon, H., Mandili, G., Medana, C., Penkman, K.E.H., Plasseraud, L., Schlichtherle, H., Taylor, S., Tokarski, C., Thomas, J., Wilson, J., Marin, F., and Demarchi, B. 2019. ‘Palaeoshellomics’ reveals the use of freshwater mother-of-pearl in prehistory. Elife 8: e45644. Crossref
Sakalauskaite, J., Marin, F., Pergolizzi, B., and Demarchi, B. 2020. Shell palaeoproteomics: First application of peptide mass fingerprinting for the rapid identification of mollusc shells in archaeology. Journal of Proteomics 227: 103920. Crossref
Schroeter, E.R. and Cleland, T.P. 2016. Glutamine deamidation: an indicator of antiquity, or preservational quality? Rapid Communications in Mass Spectrometry 30: 251–255. Crossref
Stankiewicz, B.A., Briggs, D.E.G., Evershed, R.P., Flannery, M.B., and Wuttke, M. 1997. Preservation of chitin in 25-million-year-old fossils. Science 276: 1541–1543. Crossref
Sullivan, A.P., Marciniak, S., O’Dea, A., Wake, T.A., and Perry, G.H. 2021. Modern, archaeological, and paleontological DNA analysis of a human-harvested marine gastropod (Strombus pugilis) from Caribbean Panama. Molecular Ecology Resources 21: 1517–1528. Crossref
Tan, Y.P., Hoon, S., Guerette, P.A., Wei, W., Ghadban, A., Hao, C., Miserez, A., and Waite, J.H. 2015. Infiltration of chitin by protein coacervates defines the squid beak mechanical gradient. Nature Chemical Biology 11: 488–495. Crossref
Taylor, P.D., Kudryavtsev, A.B., and Schopf, J.W. 2008. Calcite and aragonite distributions in the skeletons of bimineralic bryozoans as revealed by Raman spectroscopy. Invertebrate Biology 127: 87–97. Crossref
Thomas, P.D., Ebert, D., Muruganujan, A., Mushayahama, T., Albou, L.P., and Mi, H.Y. 2022. PANTHER: Making genome-scale phylogenetics accessible to all. Protein Science 31: 8–22. Crossref
Ward, L.W. and Andrews, G.W. 2008. Stratigraphy of the Calvert, Choptank, and St. Marys Formations (Miocene) in the Chesapeake Bay area, Maryland and Virginia. 170 pp. Virginia Museum of Natural History, Maryland.
Ward, L.W. and Gilinsky, N.L. 1988. Ecphora (Gastropoda: Muricidae) from the Chesapeake Group of Maryland and Virginia. Notulae Naturae 469: 1–21.
Welker, F., Collins, M.J., Thomas, J.A., Wadsley, M., Brace, S., Cappellini, E., Turvey, S.T., Reguero, M., Gelfo, J.N., Kramarz, A., Burger, J., Thomas-Oates, J., Ashford, D.A., Ashton, P.D., Rowsell, K., Porter, D.M., Kessler, B., Fischer, R., Baessmann, C., Kaspar, S., Olsen, J.V., Kiley, P., Elliott, J.A., Kelstrup, C.D., Mullin, V., Hofreiter, M., Willerslev, E., Hublin, J.-J., Orlando, L., Barnes, I., and MacPhee, R.D.E. 2015. Ancient proteins resolve the evolutionary history of Darwin’s South American ungulates. Nature 522: 81–84. Crossref
Welker, F., Ramos-Madrigal, J., Gutenbrunner, P., Mackie, M., Tiwary, S., Rakownikow Jersie-Christensen, R., Chiva, C., Dickinson, M.R., Kuhlwilm, M., de Manuel, M., Gelabert, P., Martinón-Torres, M., Margvelashvili, A., Arsuaga, J.L., Carbonell, E., Marques-Bonet, T., Penkman, K., Sabidó, E., Cox, J., Olsen, J.V., Lordkipanidze, D., Racimo, F., Lalueza-Fox, C., Bermúdez de Castro, J.M., Willerslev, E., and Cappellini, E. 2020. The dental proteome of Homo antecessor. Nature 580: 235–238. Crossref
Welker, F., Ramos-Madrigal, J., Kuhlwilm, M., Liao, W., Gutenbrunner, P., de Manuel, M., Samodova, D., Mackie, M., Allentoft, M.E., Bacon, A.M., Collins, M.J., Cox, J., Lalueza-Fox, C., Olsen, J.V., Demeter, F., Wang, W., Marques-Bonet, T., and Cappellini, E. 2019. Enamel proteome shows that Gigantopithecus was an early diverging pongine. Nature 576: 262–265. Crossref
Acta Palaeontol. Pol. 70 (3): 507–516, 2025
https://doi.org/10.4202/app.01201.2024